Modern Imaging Format Support

Fileglancer automatically detects modern imaging formats (OME-Zarr and N5) when you navigate into a compatible directory.
Zarr / OME-Zarr Support
Section titled “Zarr / OME-Zarr Support”Zarr preview panel
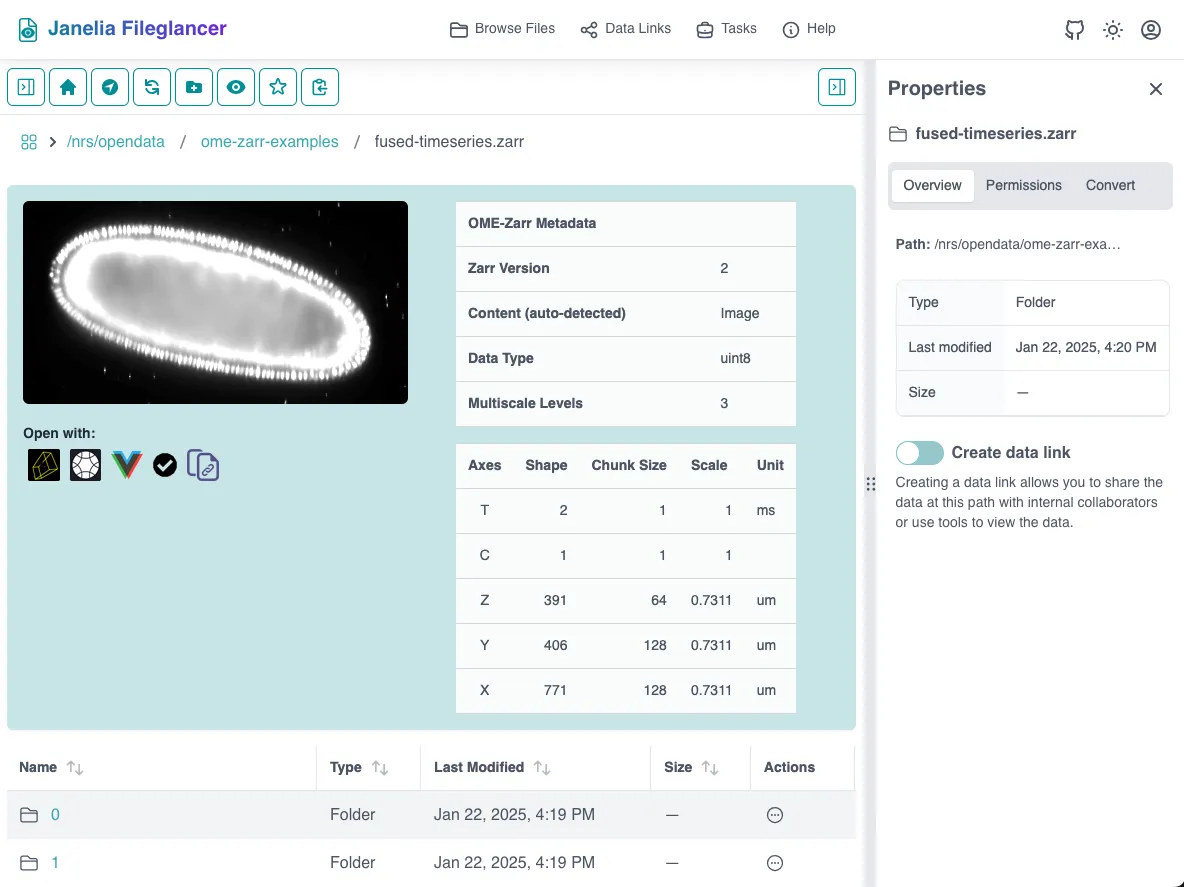
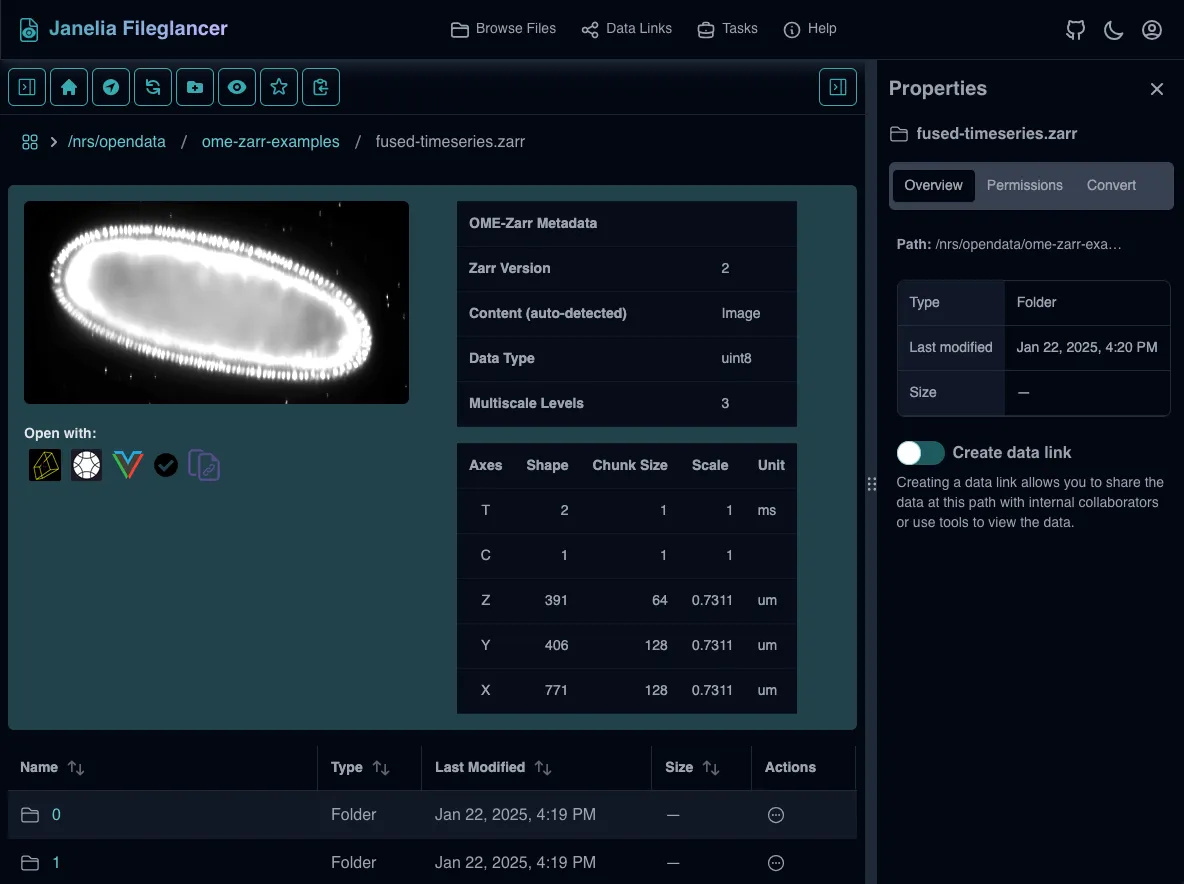
Section titled “Zarr preview panel”The Zarr preview panel shows a metadata table, thumbnail preview, and “Open with” buttons for compatible viewers for Zarr datasets.
Thumbnail previews are only generated for OME-Zarr datasets, and only if the first multiscale level is small enough to generate a thumbnail.
The metadata table includes:
- Zarr Version (v2, v3, or both if markers for multiple versions are present)
- Content (auto-detected) — heuristical image vs. segmentation detection (can be enabled/disabled in preferences)
- Data Type (e.g., uint8, float32)
- Shape and Chunk Size (for plain arrays)
- Multiscale Levels (for OME-Zarr)
- Labels (if a labels sub-directory is present)
- Per-axis details: axis name, shape, chunk size, scale, and unit
When the Zarr preview panel appears
Section titled “When the Zarr preview panel appears”The Zarr preview panel appears when Fileglancer detects Zarr marker files in the current directory. The presence of these files indicates that the directory contains Zarr data:
- Zarr v2:
.zarray(plain array) or.zattrs(OME-Zarr group) - Zarr v3:
zarr.json
For the dataset to be recognized as OME-Zarr and show the full preview panel with metadata and viewer options, the following conditions must be met:
- For a Zarr v2 group, the
.zattrsfile must contain amultiscalesfield to be recognized as OME-Zarr. - For a Zarr v3 group, the
zarr.jsonfile must containattributes.ome.multiscalesto be recognized as OME-Zarr.
Zarr data links for Neuroglancer
Section titled “Zarr data links for Neuroglancer”For Neuroglancer to open a Zarr dataset, the dataset must be:s
- organized in the way Neuroglancer expects (e.g., OME-Zarr or plain Zarr array with appropriate metadata), and
- accessible via an HTTP URL.
In Fileglancer, this means creating a data link for the directory that shows a Zarr or N5 preview panel. A data link is a proxied URL that makes the dataset accessible over the network.
- If a data link already exists, the Neuroglancer button is immediately active.
- If no data link exists, clicking the Neuroglancer button prompts you to create one.
OME-Zarr datasets (with multiscales) support all available viewers: Neuroglancer, Vol-E, Avivator (v2 only), and the OME-ZARR Validator.
Plain Zarr arrays (without multiscales) only support Neuroglancer.
Zarr data links for other viewers
Section titled “Zarr data links for other viewers”Data links can also be used to open OME-Zarr datasets in other compatible viewers. Viewers that are integrated into Fileglancer are detailed in Integrated Data Tools below, but data links can be used with any viewer that supports loading data via an HTTP URL.
N5 Support
Section titled “N5 Support”N5 metadata table
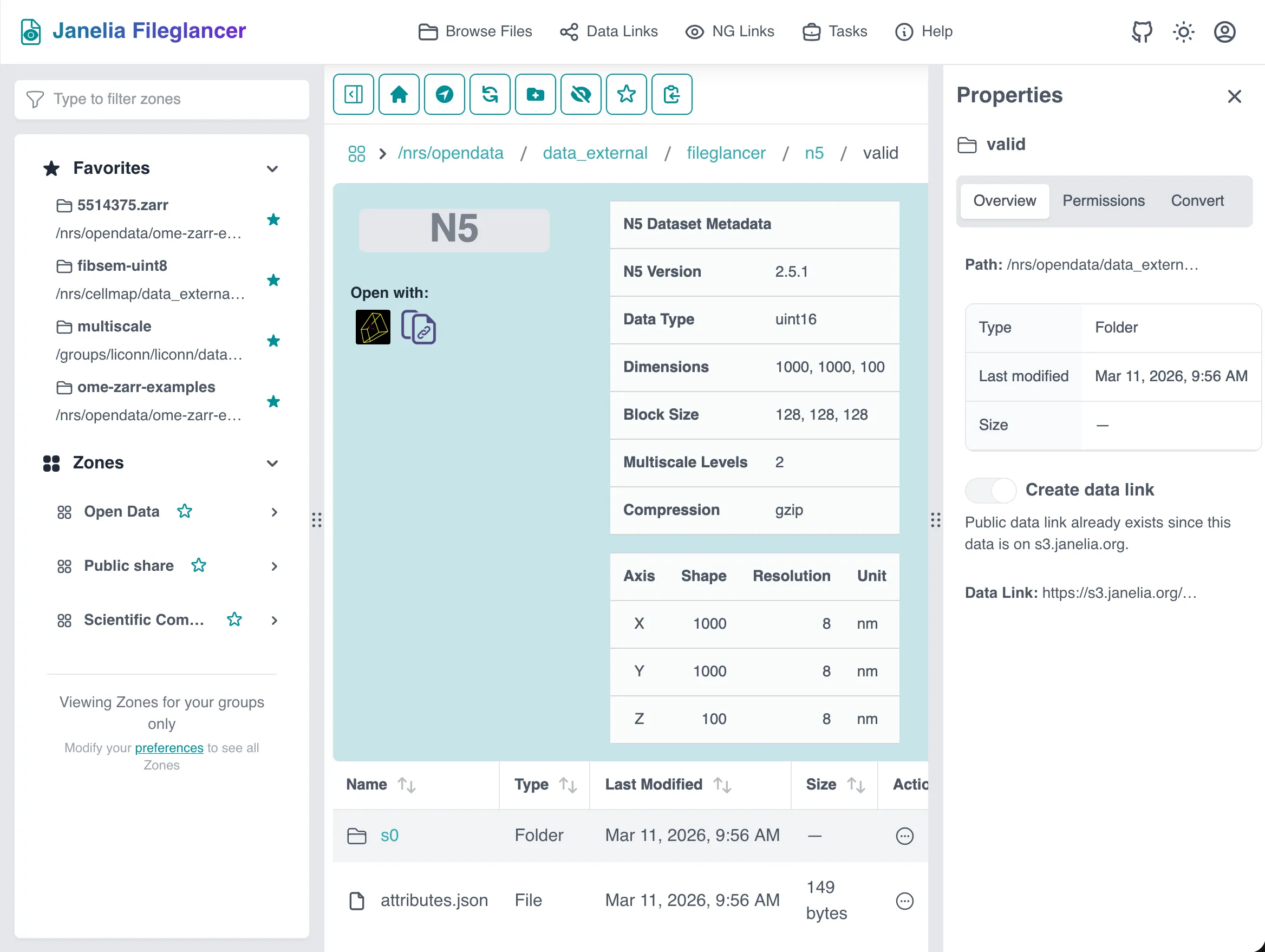
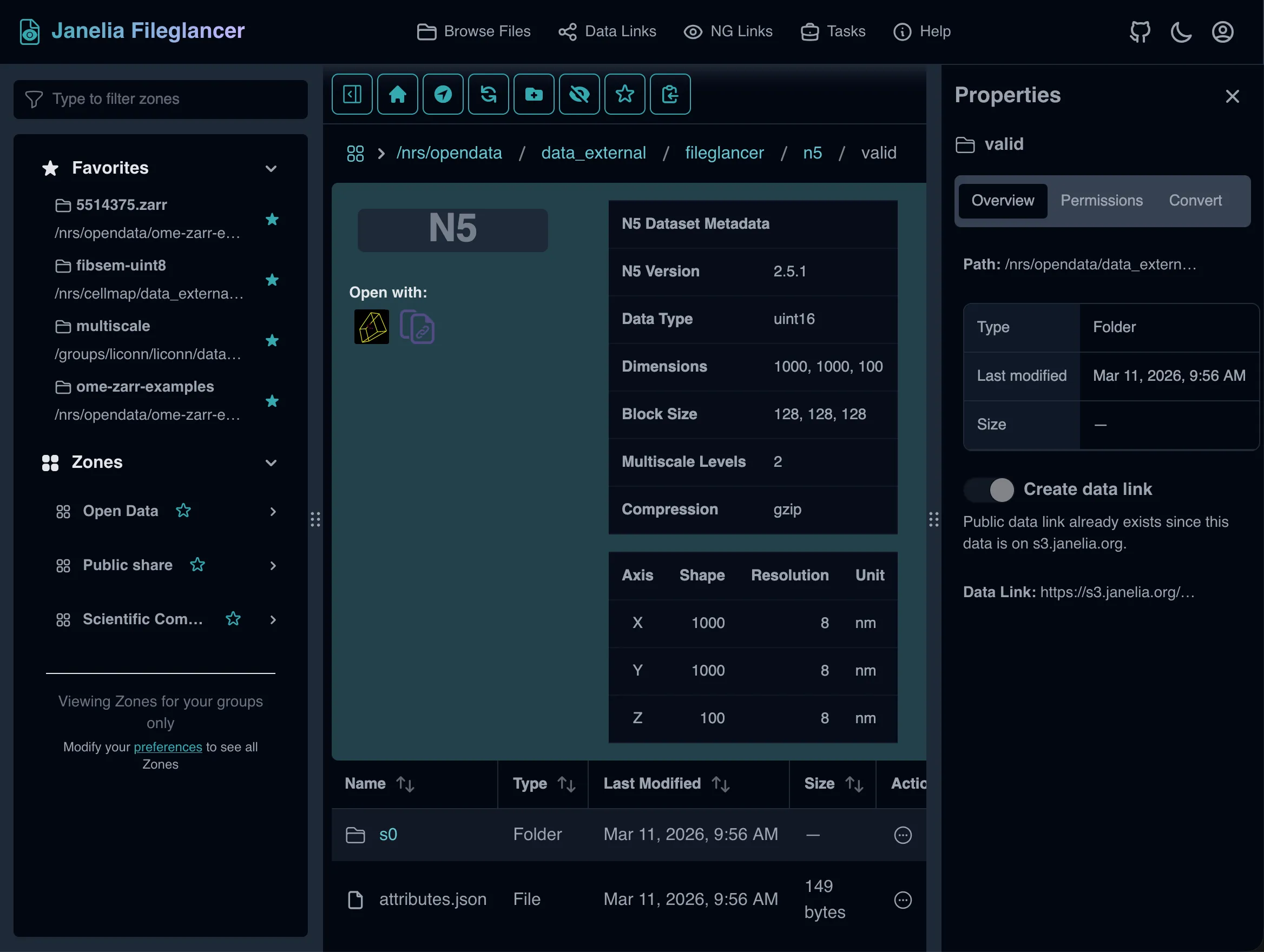
Section titled “N5 metadata table”The N5 preview panel reads metadata from attributes.json (root) and s0/attributes.json (first scale level) and displays:
- N5 Version (from the
n5field inattributes.json) - Data Type (from
s0/attributes.json, e.g., uint16) - Dimensions (from
s0/attributes.json) - Block Size (from
s0/attributes.json) - Multiscale Levels (from
downsamplingFactorsorscalesinattributes.json) - Compression (type and level, from
s0/attributes.json) - Per-axis details: axis name, shape, resolution, and unit
When the N5 preview panel appears
Section titled “When the N5 preview panel appears”Fileglancer shows the N5 preview panel when it detects both of these inside the current directory:
- An
attributes.jsonfile (the root N5 metadata) - An
s0/subdirectory (the first scale level)
Missing metadata warnings
Section titled “Missing metadata warnings”The N5 metadata table highlights missing fields that affect how viewers interpret the data:
- No resolution: If neither
resolutionnorpixelResolution.dimensionsis found inattributes.json, resolution is shown as “Unknown” with a warning: “No ‘resolution’ or ‘pixelResolution.dimensions’ found in attributes.json. Neuroglancer uses these to define the physical scale of each dimension.” - No units: If neither
unitsnorpixelResolution.unitis found inattributes.json, the unit column is blank with a warning: “No ‘units’ or ‘pixelResolution.unit’ found in attributes.json. Viewers may assume a default unit when none is specified.” - No downsampling factors: If neither
downsamplingFactorsnorscalesis found inattributes.json, multiscale levels shows “Not specified” with a warning: “No ‘downsamplingFactors’ or ‘scales’ found in attributes.json. Neuroglancer can also read ‘downsamplingFactors’ from individual sN/attributes.json files.”
N5 data links for Neuroglancer
Section titled “N5 data links for Neuroglancer”Like Zarr, N5 datasets require a data link for the directory that shows an N5 preview panel. Neuroglancer is the only integrated viewer for N5 datasets, but data links can be used with other viewers that support loading data via a HTTP URL.
Integrated Data Tools
Section titled “Integrated Data Tools”Several web-based viewers and tools for OME-Zarr and N5 datasets are directly integrated into Fileglancer. When you open a compatible dataset, “Open with” buttons appear in the preview panel to launch these tools directly from Fileglancer.
- A powerful web-based viewer for large-scale volumetric data.
- Supported formats: OME-Zarr, plain Zarr arrays, N5
- Allen Cell Institute’s volume viewer designed for cellular and subcellular imaging.
- Supported formats: OME-Zarr only
- A lightweight, web-based image viewer for OME-Zarr data.
- Supported formats: OME-Zarr v2 only
- A diagnostic tool for verifying OME-Zarr datasets against the OME-Zarr specification.
- Supported formats: OME-Zarr only
Accessing Data Tools from Fileglancer
Section titled “Accessing Data Tools from Fileglancer”Locating Tool Options
Section titled “Locating Tool Options”When you navigate to a compatible dataset (OME-Zarr, Zarr, or N5 format):
-
View the file/directory in Fileglancer
Navigate to your dataset directory. The dataset will be recognized automatically if it meets the Zarr or N5 criteria, and the preview panel will appear above the file list. -
Find the “Open with” section
Located in the file preview area, below the thumbnail preview of the dataset, there is a list of compatible viewers with their respective logos. There is also a button to copy the direct data link.
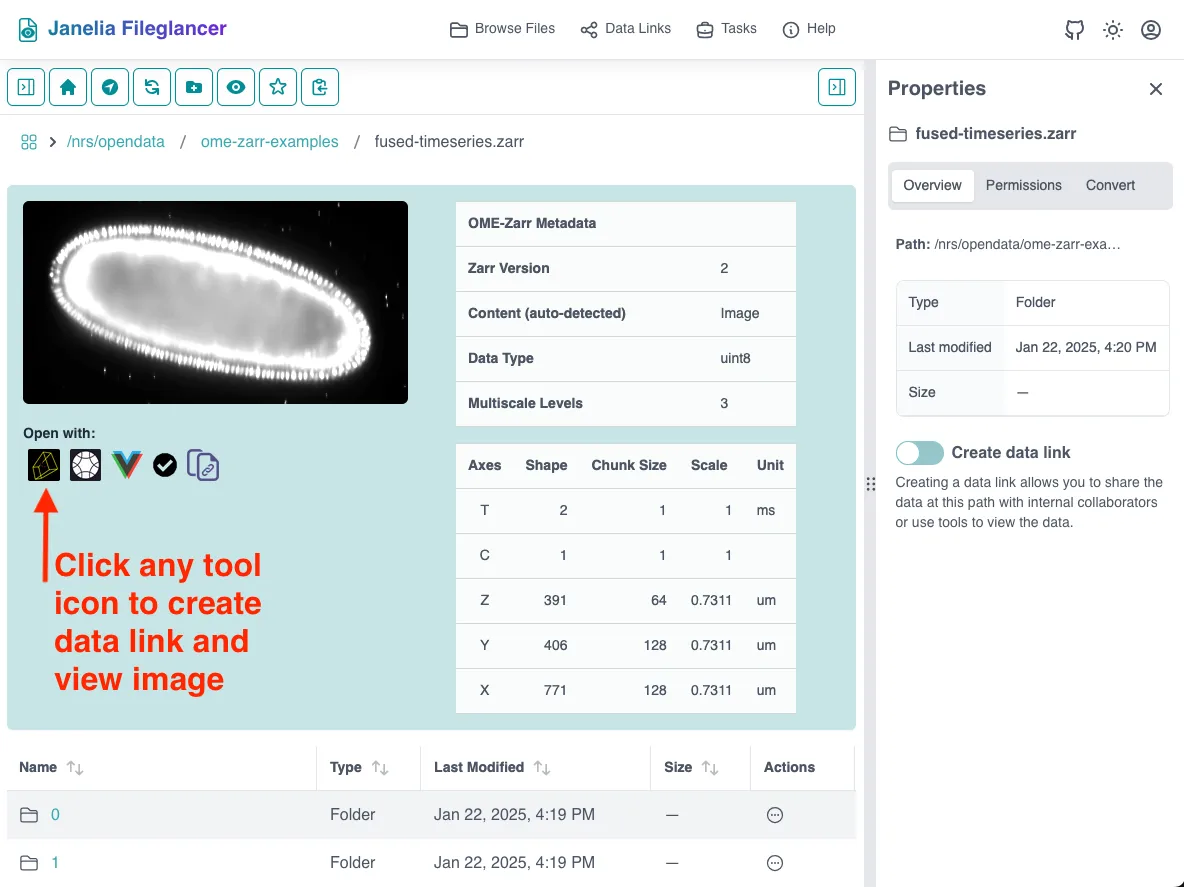
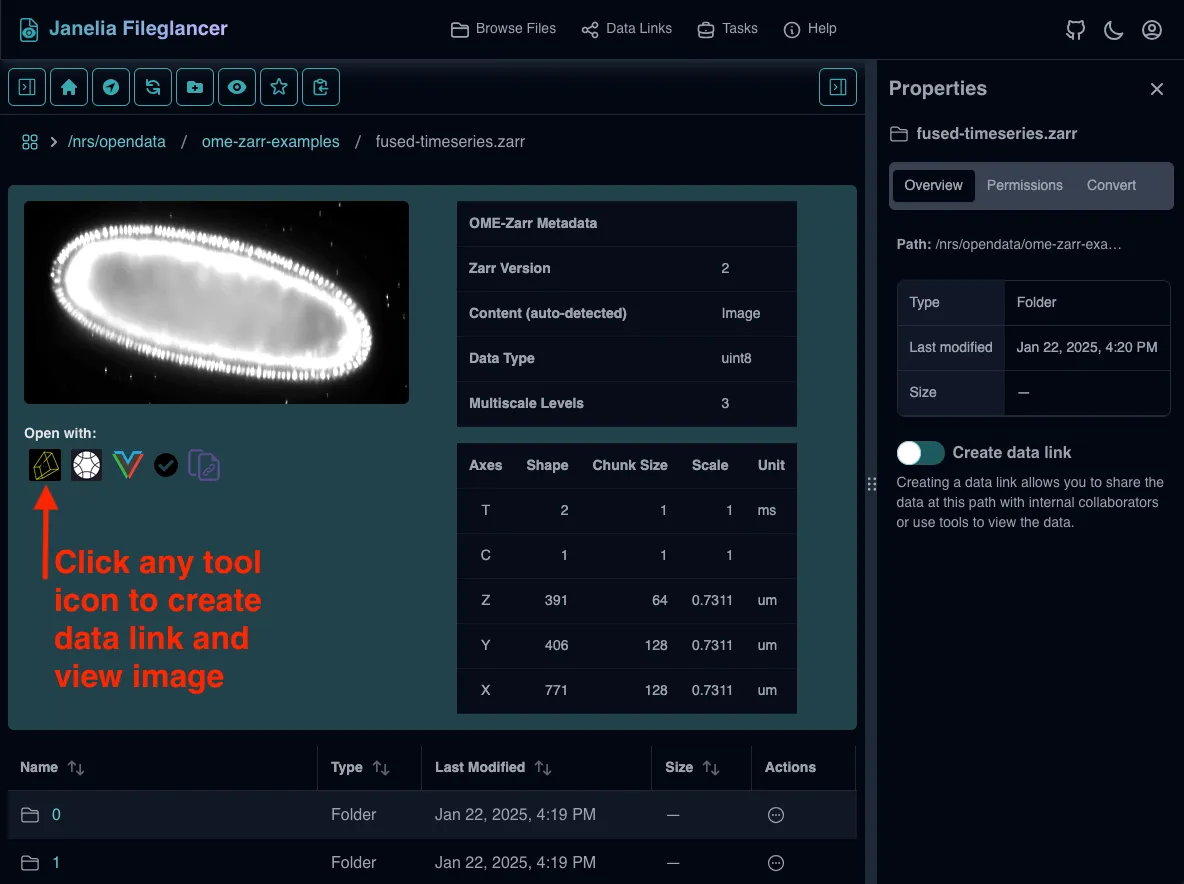
Launching Viewers
Section titled “Launching Viewers”-
Click the desired viewer button Each viewer opens in a new browser tab once a data link is available.
-
Share viewer URLs Copy the URL from the opened viewer tab. Share with internal collaborators. URLs remain valid as long as data links are active.
Troubleshooting
Section titled “Troubleshooting”Preview Panel Not Appearing
Section titled “Preview Panel Not Appearing”For Zarr directories:
- Verify the directory contains at least one Zarr marker file:
zarr.json,.zarray, or.zattrs - For OME-Zarr, ensure the metadata includes a
multiscalesfield
For N5 directories:
- Verify the directory contains both
attributes.jsonand ans0/subdirectory - Ensure
s0/attributes.jsonexists and is readable
Data Loading Issues
Section titled “Data Loading Issues”Problem: A tool opens but no data appears
- Verify data format compatibility
- Ensure data structure is correct
- Request conversion to compatible format
Performance Issues
Section titled “Performance Issues”Problem: Slow data loading
- Check data chunk sizes and compression
- Consider creating optimized versions for viewing
- Verify network connectivity and speed
Sharing Issues
Section titled “Sharing Issues”Problem: Shared URLs don’t work for collaborators
- Check the data link is still active
- Ensure collaborators are on correct network